As discussed in a previous post, in order for the program to work using the new CNCPS 6.5 biology, users must first convert farms using the Amino Acid Conversion Tool. A quick video demonstrating that process is found here. There are a couple of issues about how that task is actualized that deserve repeating, or, in some cases, highlighting.
-
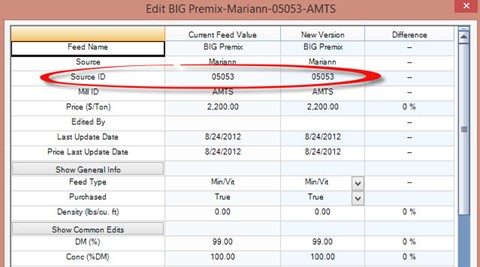
The program will match the farm feed to an AMTS Feedbank feed based on the farm feed’s original source code.

This means, the amino acid profile of the farm feed will be dependent on the amino acids profile of the AMTS Feedbank feed chosen when the farm feeds were set up. Which brings us to:
-
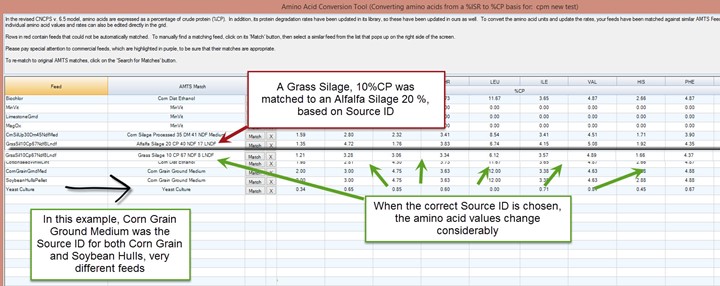
The Source ID is very important. In some cases it can be a little wonky. Those are;
- When it is a commercial protein or amino acid source that you roughly matched to a library feed in previous versions of the program. Those profiles may be very different. Check with the amino acid company for the correct values. Those can be inputted directly in the Amino Acid conversion Tool Screen.
- When your farms originated as a CPM file. Those feed libraries were not as extensive as the CNCPS library (which is the base for the AMTS library) and many feeds shared the same Source ID. The conversions can then be pretty inaccurate
-
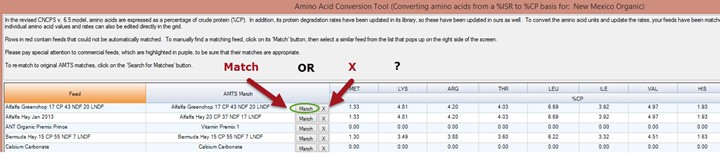
Prior to conversion, some users have modified the farm feed amino acid profile from that of the library feed Source ID. Usually this is because the feed has been analyzed by lab and the values are specific to that farm feed, not an average of many feeds, as library values are. In this case, the user would not like to lose those values. We have a solution for that!

The Match button is used when the user wants to convert the amino acid values to the AMTS Feedbank values (those shown). The X button is used when the user has altered the amino acid values in the farm file and wants to maintain those numbers. When the X is clicked, the values inputted will show in the amino acid value columns.